# Packages
library(sf)
library(dplyr)
library(ggplot2)
library(gstat)
library(terra)
library(spatstat.geom)
library(spatstat.explore)
sf_use_s2(FALSE)
# Data
data_path <- "C:/Users/celio/Inter Dens/Data"
luft <- st_read(file.path(data_path, "luftqualitaet.gpkg"), layer = "luftqualitaet", quiet = TRUE)
schweiz <- st_read(file.path(data_path, "schweiz.gpkg"), layer = "schweiz", quiet = TRUE)
milan <- st_read(file.path(data_path, "rotmilan.gpkg"), layer = "rotmilan", quiet = TRUE)
luft <- st_transform(luft, 2056)
schweiz <- st_transform(schweiz, 2056)
milan <- st_transform(milan, 2056)Interpolation and Density Estimation
Interpolation and Density Estimation
This document contains the solutions for interpolation (NO₂) and density estimation (red kite data).
#Setup
Interpolation Exercise 1: IDW
grid <- st_make_grid(schweiz, cellsize = 10000, what = "centers") |>
st_sf() |>
st_intersection(schweiz)
idw_res <- gstat::idw(
value ~ 1,
luft,
newdata = grid,
idp = 2,
nmin = 4,
nmax = 14
)[inverse distance weighted interpolation]ggplot() +
geom_sf(data = idw_res, aes(color = var1.pred), size = 1.5) +
geom_sf(data = schweiz, fill = NA, color = "black") +
geom_sf(data = luft, color = "red", size = 1) +
scale_color_viridis_c(name = expression(NO[2])) +
labs(title = "IDW interpolation of NO2") +
theme_minimal()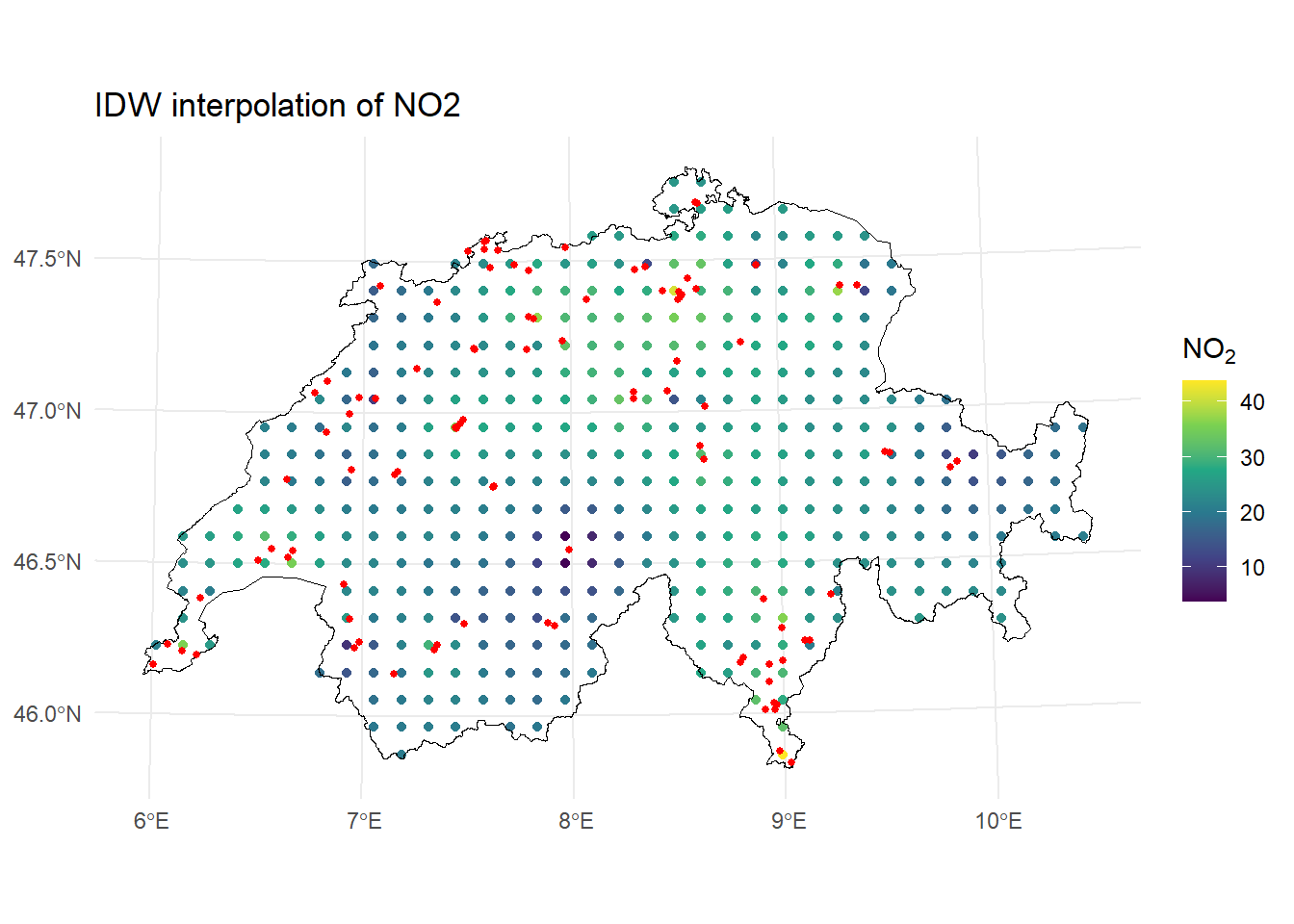
# IDW → raster
idw_vect <- vect(idw_res)
idw_r <- rast(ext(idw_vect), resolution = 10000, crs = st_crs(schweiz)$wkt)
idw_r <- rasterize(idw_vect, idw_r, field = "var1.pred")
plot(idw_r, main = "IDW interpolation raster")
plot(vect(schweiz), add = TRUE)
points(st_coordinates(luft), pch = 16, cex = 0.5)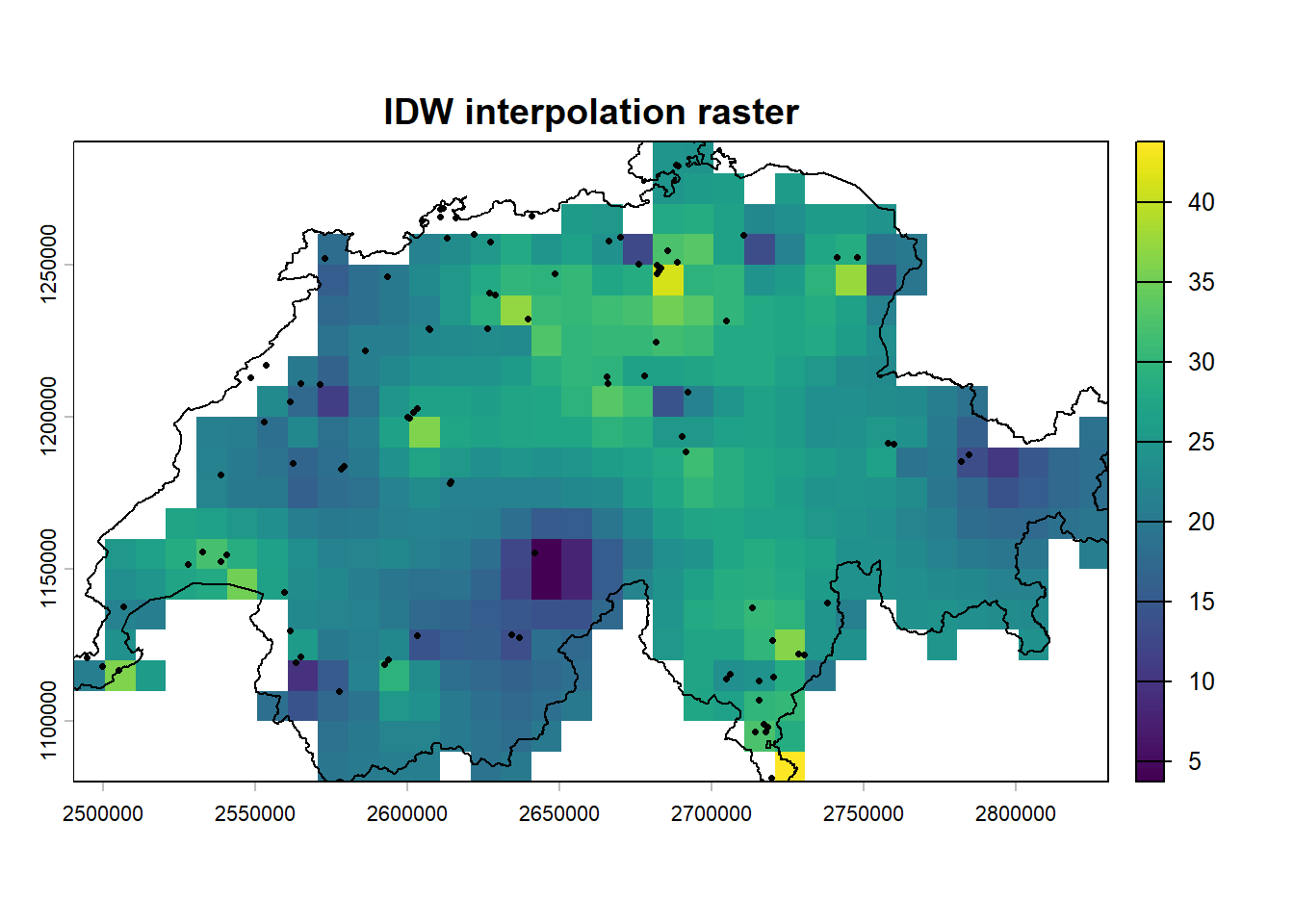
Interpolation Exercise 2: Nearest Neighbour
vor <- st_voronoi(st_union(luft)) |>
st_collection_extract("POLYGON") |>
st_as_sf()
vor <- st_join(vor, luft) |>
st_intersection(schweiz)
ggplot() +
geom_sf(data = vor, aes(fill = value), color = NA) +
geom_sf(data = schweiz, fill = NA, color = "black") +
geom_sf(data = luft, color = "red", size = 1) +
scale_fill_viridis_c(name = expression(NO[2])) +
labs(title = "Nearest neighbour (Voronoi)") +
theme_minimal()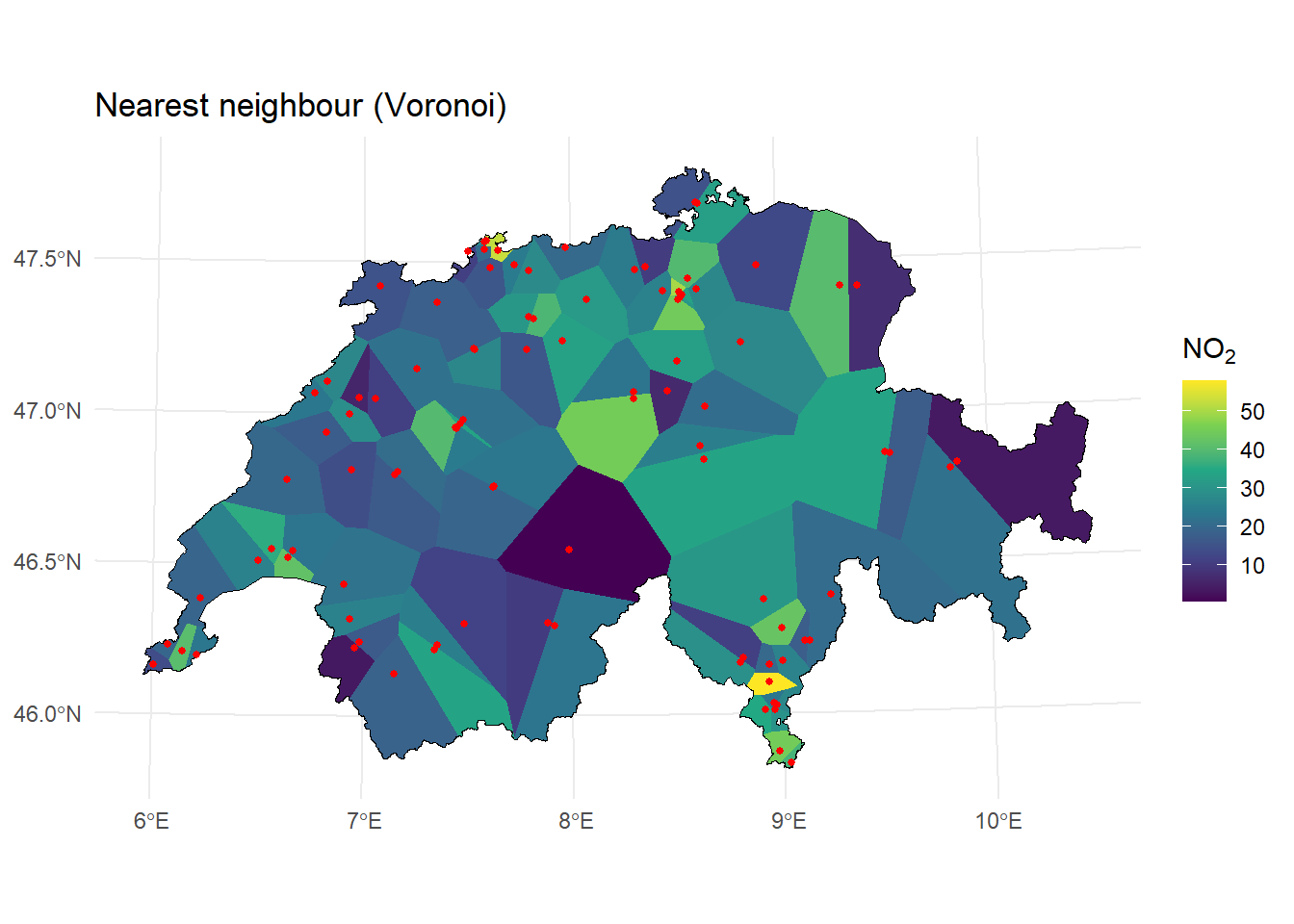
Density Estimation Exercise 1: Kernel Density Estimation
pp <- as.ppp(st_geometry(milan))
bw.diggle(pp) sigma
25.16005 bw.CvL(pp) sigma
38038.72 bw.scott(pp) sigma.x sigma.y
4302.234 1829.652 bw.ppl(pp) sigma
944.463 sigma <- c(5000, 5000)
kde <- density(pp, sigma = sigma, eps = 500)
kde_r <- rast(kde)
plot(kde_r, main = "Kernel density estimation")
points(pp, col = "grey", pch = 16, cex = 0.2)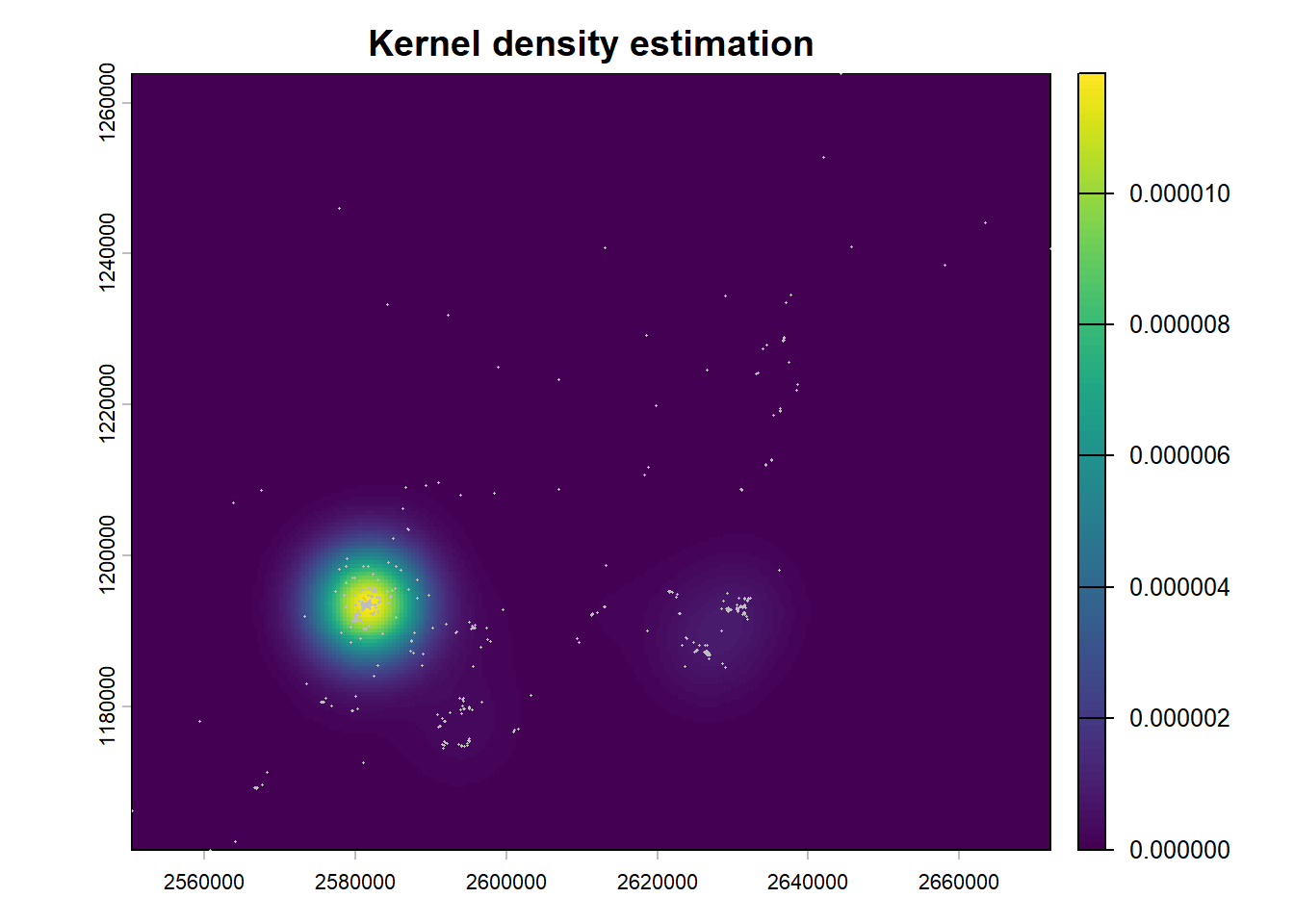
Density Estimation Exercise 2: Voronoi
# =========================
# VORONOI (POINT STRUCTURE)
# =========================
vor_m <- st_voronoi(st_union(milan)) |>
st_collection_extract("POLYGON") |>
st_sf()
vor_m <- st_intersection(vor_m, schweiz)
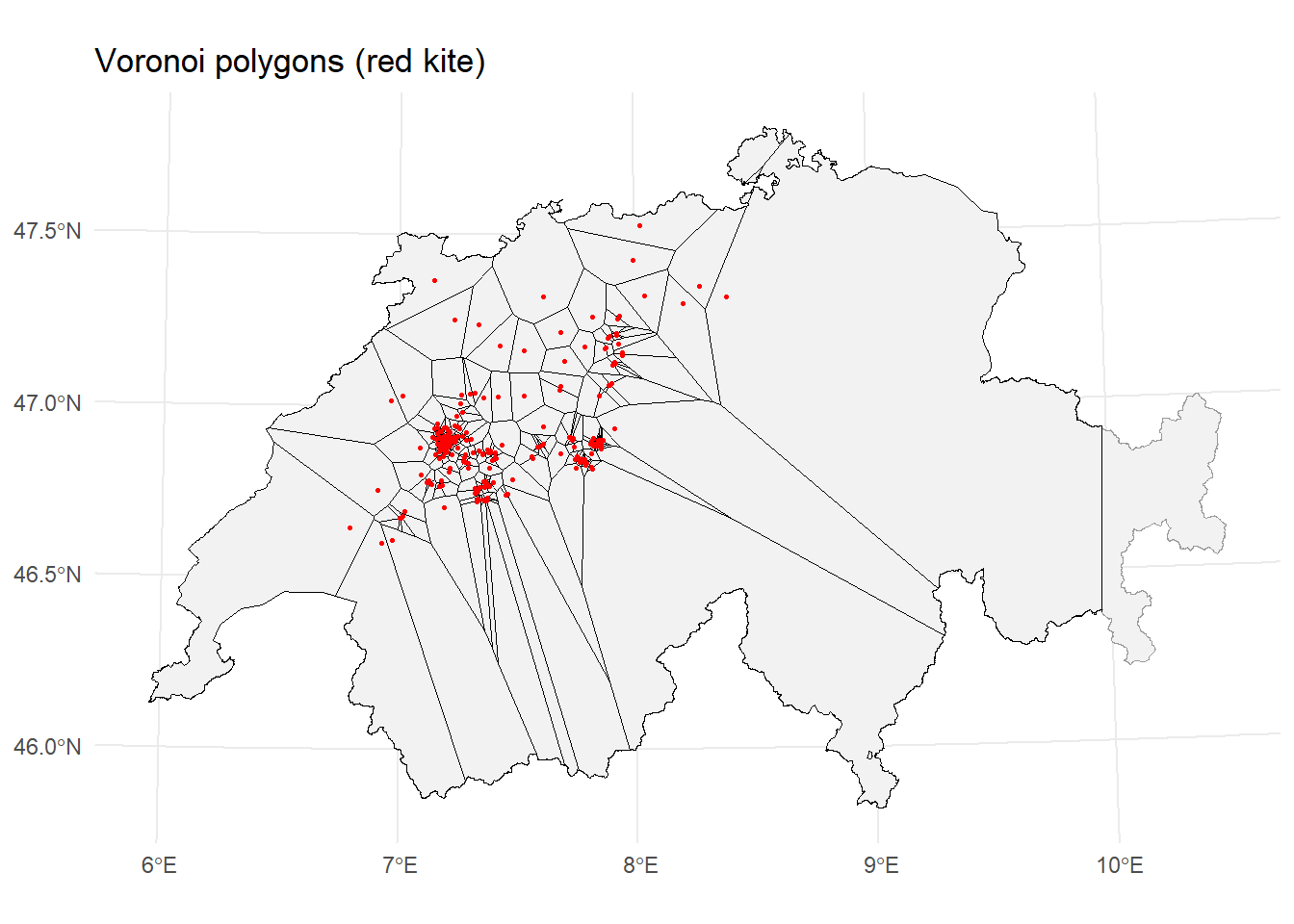
ggplot() +
geom_sf(data = schweiz, fill = "grey95", color = "grey60") +
geom_sf(data = vor_m, fill = NA, color = "black", linewidth = 0.2) +
geom_sf(data = milan, color = "red", size = 0.5) +
labs(title = "Voronoi polygons (red kite)") +
theme_minimal()